Why Would You Biotinylate Ubiquitin?
E-STUB: A Precision Tool for Mapping E3 Ligase Substrates
In march 2025, colleagues from Dana Faber and The Broad Institute published in 𝘕𝘢𝘵𝘶𝘳𝘦 𝘊𝘩𝘦𝘮𝘪𝘤𝘢𝘭 𝘉𝘪𝘰𝘭𝘰𝘨𝘺:
“𝘜𝘣𝘪𝘲𝘶𝘪𝘵𝘪𝘯-𝘴𝘱𝘦𝘤𝘪𝘧𝘪𝘤 𝘱𝘳𝘰𝘹𝘪𝘮𝘪𝘵𝘺 𝘭𝘢𝘣𝘦𝘭𝘪𝘯𝘨 𝘧𝘰𝘳 𝘵𝘩𝘦 𝘪𝘥𝘦𝘯𝘵𝘪𝘧𝘪𝘤𝘢𝘵𝘪𝘰𝘯 𝘰𝘧 𝘌3 𝘭𝘪𝘨𝘢𝘴𝘦 𝘴𝘶𝘣𝘴𝘵𝘳𝘢𝘵𝘦𝘴”
How it works
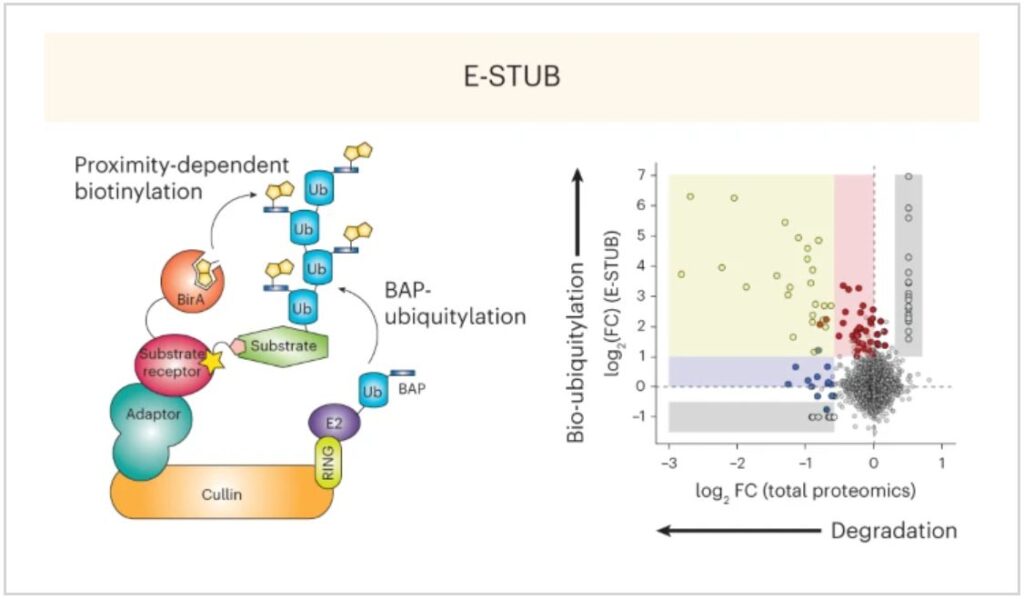
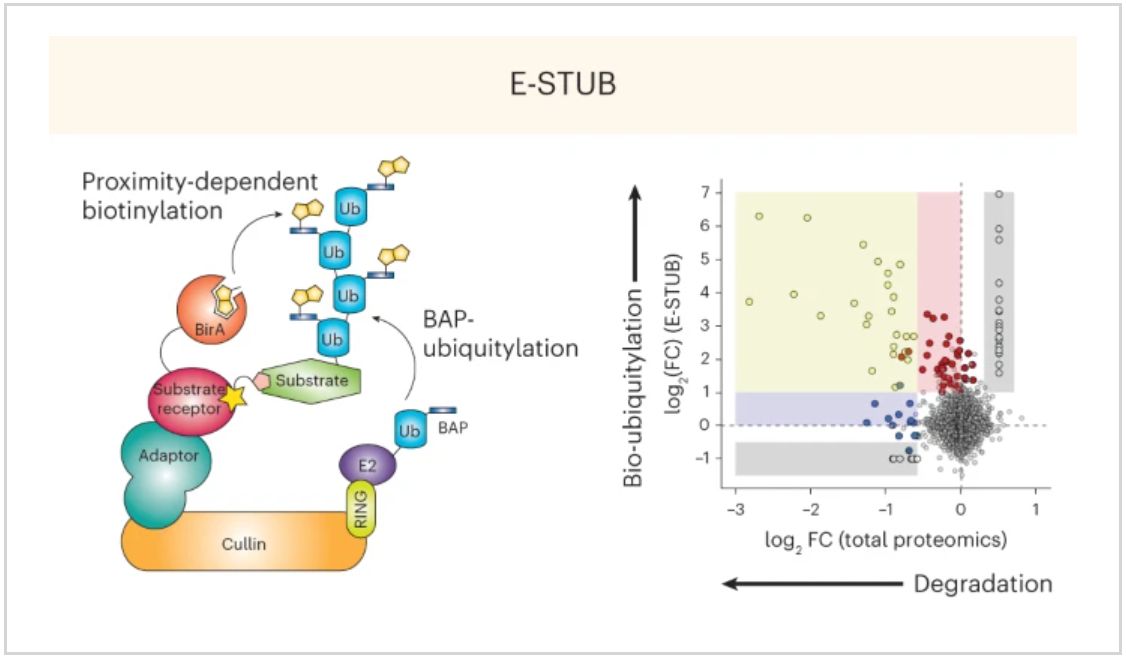
E-STUB (E3-Substrate Tagging by Ubiquitin Biotinylation) combines the principles of proximity labeling with the functional specificity of ubiquitin enrichment. The method involves “co-expression of a ubiquitin harboring a biotin acceptor peptide (BAP; for example, AviTag) and an E3 ligase of interest fused to the Escherichia coli biotin ligase BirA”, which “would result in biotinylation of ubiquitylated substrates in the proximity of a given E3 ligase”
My Highlights
✅ High specificity: Optimized A3-ubiquitin yields low background and robust signal, uncovering known targets of molecular glues (e.g., GSPT1 with CC-885) and PROTACs with kinase selectivity.
(A3 = optimized biotin acceptor peptide)
✅ Reveals ubiquitylation events that don’t always lead to degradation (consistent with previous studies)
✅ Collateral ubiquitylation mapping: Captures modification of neighbouring complex members. The authors found that “many members of corepressor complexes are significantly bio-ubiquitylated and degraded, while HDAC1/HDAC2 were bio-ubiquitylated but more resistant to degradation”
✅ Broad applicability: Validated across multiple E3 ligases (CRBN, VHL, DCAF15) and degraders, plus detection of physiological substrates like HIF1A via VHL loss-of-function profiling.
Limitations
The authors acknowledge that “head-to-head comparison between two distinct ligases generates many statistically significant hits that could be false positives due to differences in expression levels and local backgrounds” and suggest that probabilistic scoring approaches could provide better confidence estimation once more E3 ligases have been profiled.
Which E3 ligase would you like to test using E-STUB ❓
Full manuscript: https://www.nature.com/nature-index/article/10.1038/s41589-024-01590-9
Leave your comment under my LinkedIn post here.