Let it GLOW: Novel Proximity Inducing Modality for DNA Modification
DNA methyltransferases (MTases) can recognize specific DNA sequences and transfer a methyl group from the S-adenosyl-L-methionine (SAM) cofactor to a target nucleobase. This ability has been exploited for sequence-specific DNA labeling using synthetic SAM analogues (SMILing, mTAG).
However, both approaches (SMILing and mTAG) share following limitations:
🔸 The SAM analogues are chemically reactive (labile aziridine rings or sulfonium centers)
🔸 Labeling fails if the target nucleobase is already methylated by endogenous processes
𝗞𝗲𝘆 𝗶𝗻𝗻𝗼𝘃𝗮𝘁𝗶𝗼𝗻
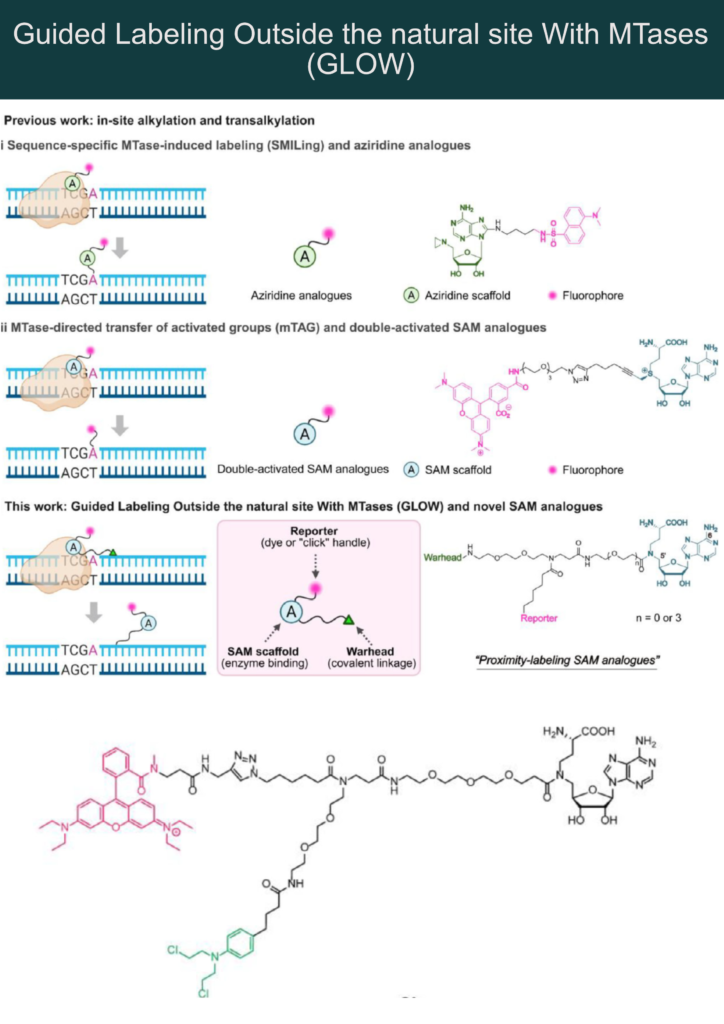
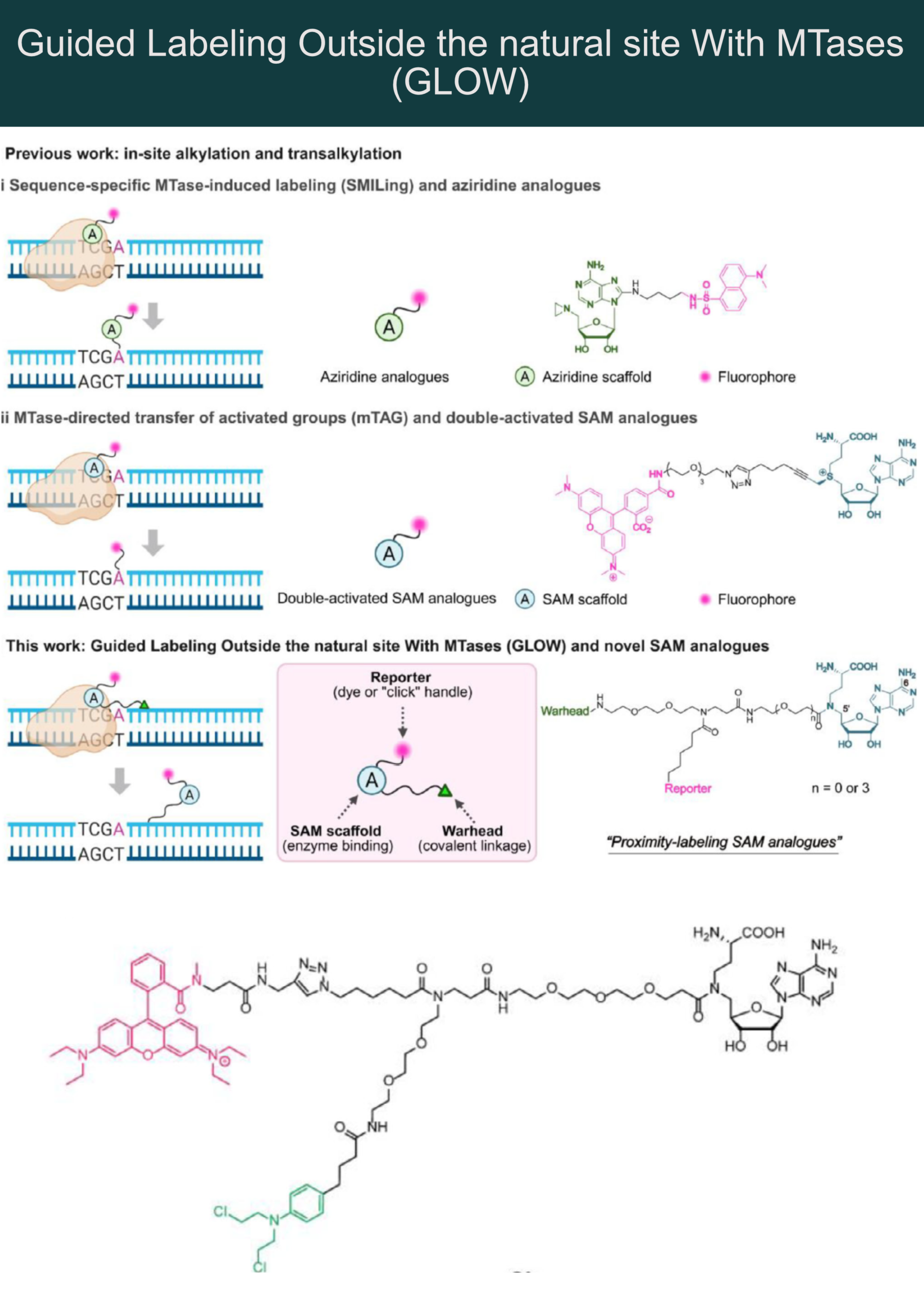
Chen et al. now report a novel proximity inducing S-adenosyl-L-methionine (SAM) mimetic modality that induces so-called “GLOW” (Guided Labeling Outside the natural site With MTases).
Instead of labeling at the target DNA nucleobase, this SAM mimetic modality uses the MTase only to induce proximity in a DNA-sequence-specific manner. This was achieved by replacing the unstable sulfonium/aziridine center with a stable amide moiety.
𝗛𝗼𝘄 𝗶𝘁 𝘄𝗼𝗿𝗸𝘀
The “GLOW modality” is tri-functional:
🔸 SAM mimicking moiety for MTase binding (in dark blue – see the figure)
🔸 Reactive warhead for covalent DNA attachment (green)
🔸 Reporter group (azide or fluorophore) for detection (pink)
1) SAM mimetic moiety binds to MTase that simultaneously binds to a specific DNA sequence
2) The reactive “warhead” covalently modifies nucleobases adjacent to the recognition site.
𝗠𝘆 𝗵𝗶𝗴𝗵𝗹𝗶𝗴𝗵𝘁𝘀
✅ Six proximity-labeling SAM analogues synthesized with psoralen or chlorambucil warheads
✅ Chlorambucil variants with C-5′ modification outperformed psoralen variants
✅ Lead analogue 2d achieved ~70% labeling efficiency on a plasmid containing 12 target sites
✅ Sequence specificity confirmed by single-molecule fluorescence consensus mapping
✅ Fluorescent analogue 3b (the molecule in the figure below) enabled direct one-step DNA staining
𝗠𝗲𝘁𝗵𝘆𝗹𝗮𝘁𝗶𝗼𝗻 𝗶𝗻𝗱𝗲𝗽𝗲𝗻𝗱𝗲𝗻𝗰𝗲
On pre-methylated DNA, a conventional double-activated SAM analogue failed to label. GLOW molecule 3b retained sequence-specific labeling, confirming that this approach bypasses interference from endogenous methylation.
This is a conceptually distinct mode of MTase-directed DNA labeling with potential for genome mapping, epigenetic profiling, and single-molecule imaging in native genomic contexts.
𝗦𝗵𝗮𝗿𝗲 𝘆𝗼𝘂𝗿 𝗼𝗽𝗶𝗻𝗶𝗼𝗻 🧐
Do you think that this concept can be adopted for development of therapeutic proximity inducing modalities?
Full paper: https://lnkd.in/g7wnF8SF