Editing Protein Backbone Inside Living Cells
For decades, creating proteins with extended backbones (β, γ, δ-amino acids) inside living cells has been challenging.
Researchers at University of California, Berkeley (Alanna Schepartz, Leah Roe), Yale University, Amgen and collaborators developed BEAR reactions to address this challenge.
𝐖𝐡𝐲 𝐝𝐨𝐞𝐬 𝐭𝐡𝐢𝐬 𝐦𝐚𝐭𝐭𝐞𝐫?
→ Extension of backbone can lead to improved phys-chem properties
→ Better stability and resistance to enzymes
→ Enhanced cellular uptake for therapeutic applications
→ Altered recognition by immune system
Traditional protein ingeneering methods often fails because:
❌ Ribosomes struggle with non-natural monomers
❌ EF-Tu can’t deliver the right acylated tRNAs
❌ The cellular machinery just wasn’t designed for this
(EF-Tu = Elongation Factor Tu – a bacterial protein essential for protein synthesis)
𝐒𝐨𝐥𝐮𝐭𝐢𝐨𝐧:
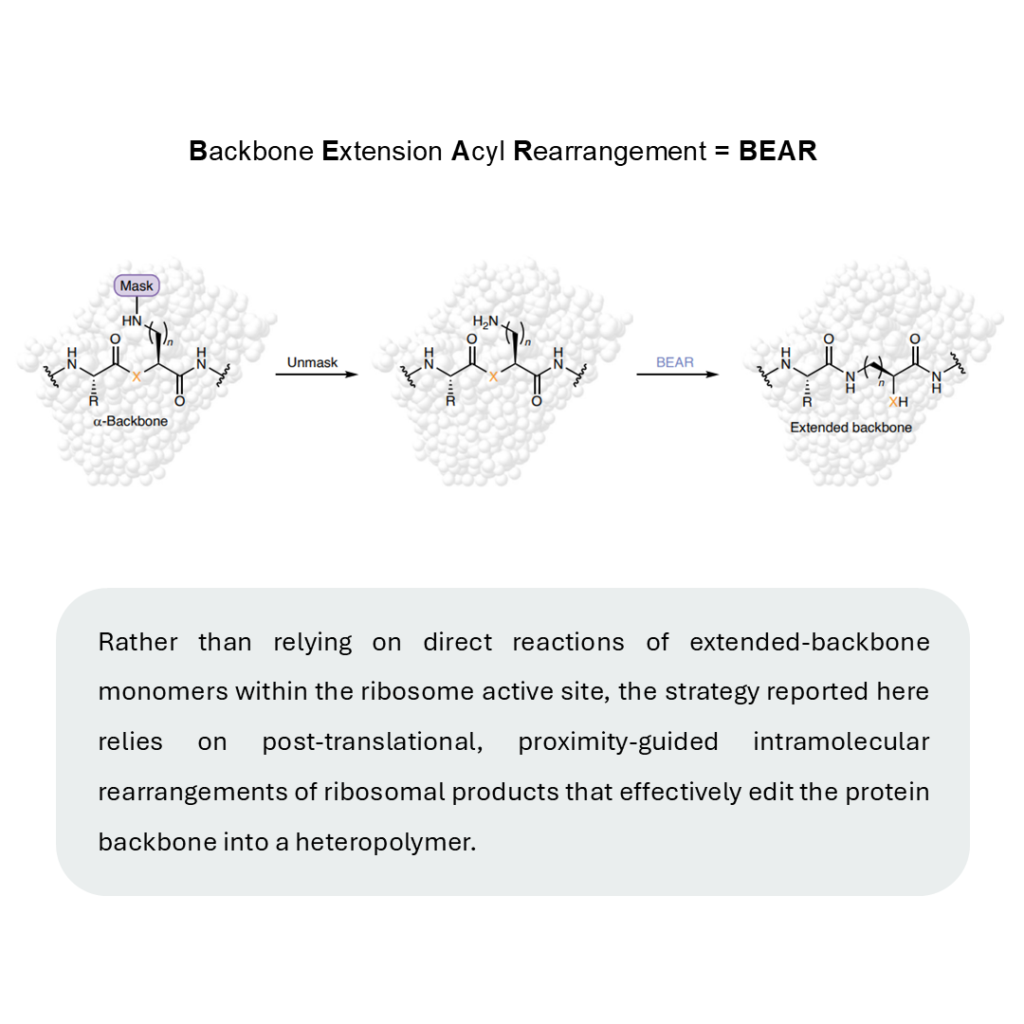
BEAR reactions (Backbone Extension Acyl Rearrangements).
Instead of forcing cells to directly make extended proteins, researchers developed a clever workaround:
1️⃣ Incorporate masked precursors using existing cellular machinery
2️⃣ Unmask the nucleophiles post-translationally
3️⃣ Rearrange spontaneously via thermodynamically favored reactions
Results:
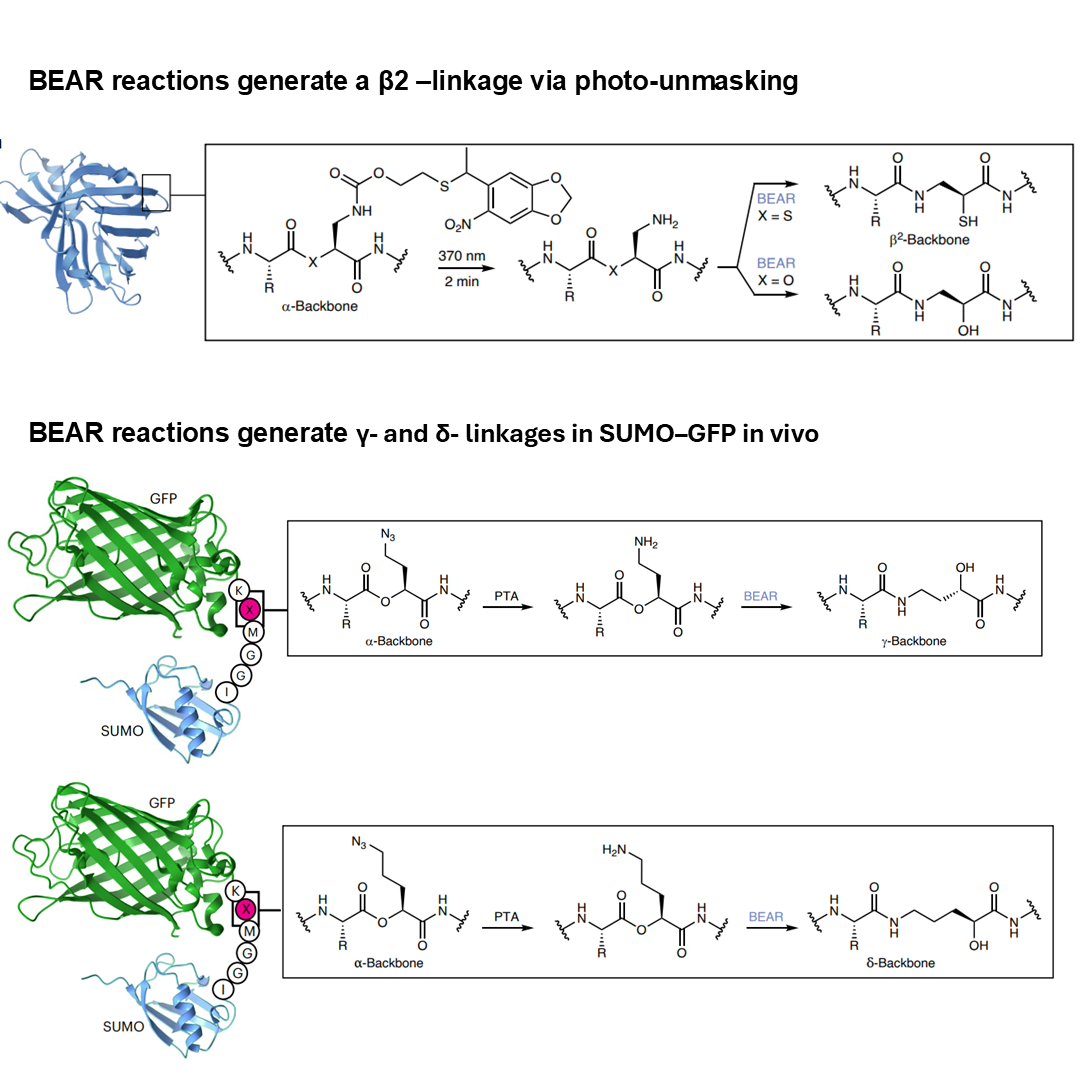
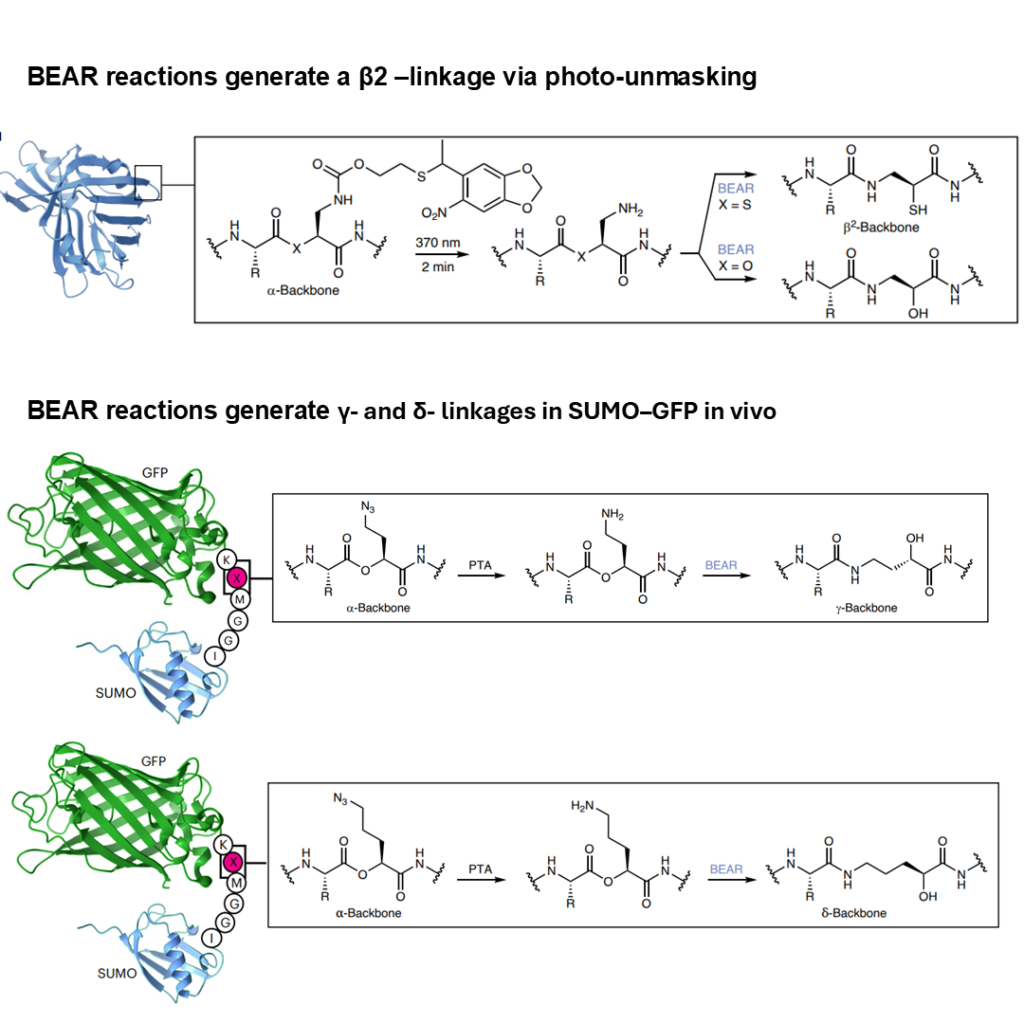
✅ β₂-linkages: Successfully demonstrated in NanoLuc
✅ γ-linkages: >90% efficiency with optimized conditions
✅ δ-linkages: 27% yield (challenging but achievable)
𝐓𝐡𝐢𝐬 𝐨𝐩𝐞𝐧𝐬 𝐭𝐡𝐞 𝐝𝐨𝐨𝐫 𝐭𝐨:
✔️ Next-generation protein therapeutics
✔️ Enhanced biomaterials
✔️ Evolution of extended-backbone protein hybrids
What I like the most is that this protocol relies only on established activities of known orthogonal enzymes and the wild-type E. coli ribosome” and “does not require engineering of EF-Tu, tRNA or the ribosome”
Where do you see the biggest potential of this methodology❓
Full manuscript: https://pubmed.ncbi.nlm.nih.gov/40841495/
Leave your comments under my LinkedIn post here.
#Biotech #ProteinEngineering #ChemicalBiology #Innovation #DrugDiscovery #Biochemistry #ProteinBiochemistry