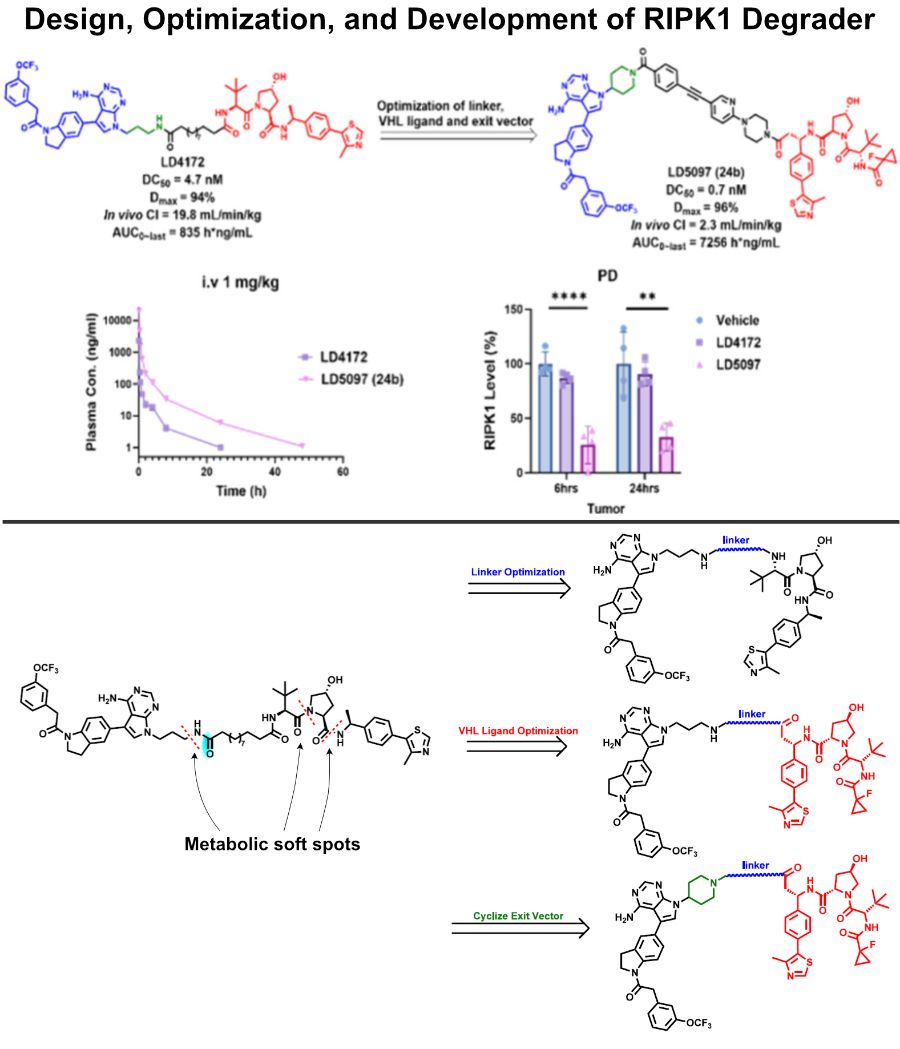
Design, Development and Optimization of RIPK1 Degrader
Optimization of PROTACs for in vivo applications is non-trivial, thought not impossible.
A nice example on this topic represents freshly published paper in J. Med. Chem., showcasing successful optimization of a RIPK1 PROTAC degrader.
Previous RIPK1 degrader LD4172 suffered from metabolic degradation and fast clearance (I highlighted the metabolic soft spots in the figure below).
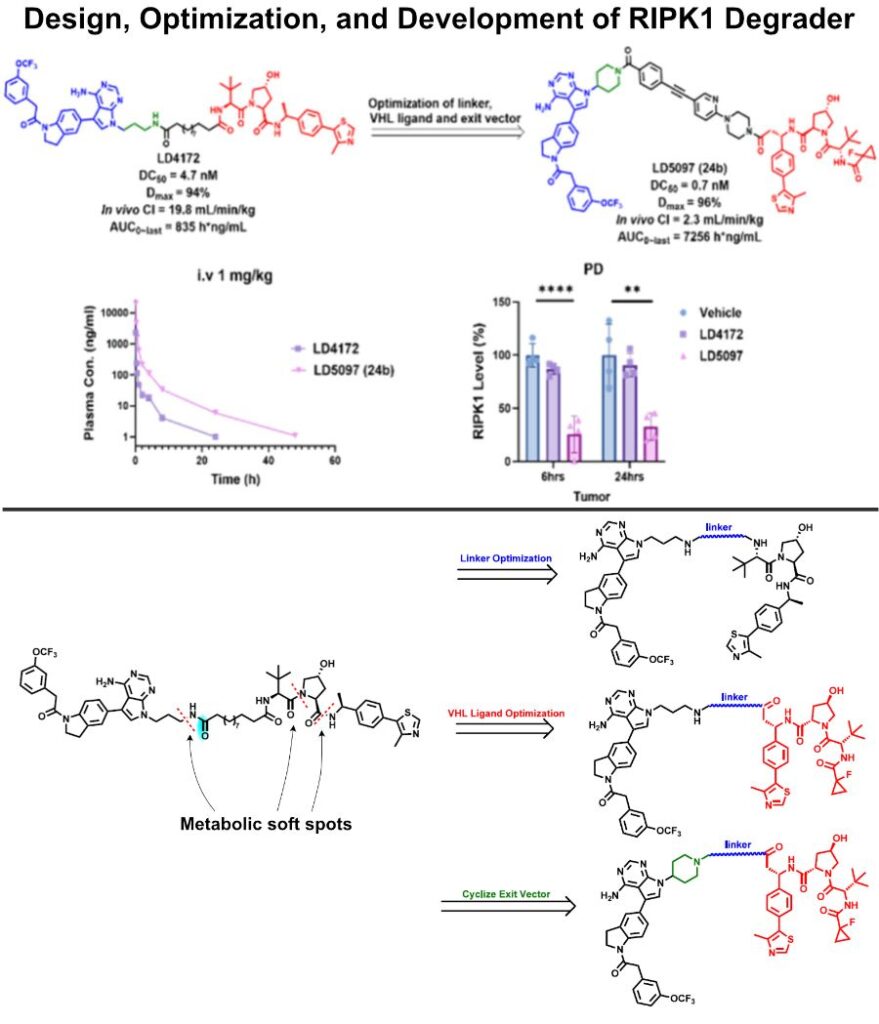
✅ To improve the PK profile, researchers subjected LD4172 to systematic medicinal chemistry optimization:
– RIPK1 linker attachment optimization: rigid piperidine
– VHL ligand with alternative exit vector performed the best
– Screening of rigid and flexible linkers
✅ This resulted in LD5097 that demonstrates:
– Potent RIPK1 degradation across multiple cancer cell lines (DC50: 0.8-2.8 nM)
– High selectivity for RIPK1: no off-target observed by MS-proteomics (17,000+ proteins analyzed)
– Excellent plasma half-life: 7.3 hours
– Reduced clearance (2.3 mL/min/kg).
– Superior tumor penetration: A single dose of LD5097 at 10 mg/kg achieved >70% RIPK1 degradation in mouse xenograft model.
💡 The SAR shows that most linkers led to loss of activity, showing that extensive linker screening is often needed in PROTAC development. Especially for rigid linker where the SAR can be very steep.
💡 Species-Dependent Activity
Interestingly, LD5097 exhibited significantly greater activity in human cell lines compared to murine cell lines (>100-fold difference), attributed to differences between human and mouse RIPK1 and potential variations in ternary complex geometry, highlighting a critical translational consideration for preclinical development.
💡 Let me know your thoughts
How would you approach the medchem optimization? Would you do anything differently?
Have you encountered species-dependent activity in your projects?
The full research paper: https://pubs.acs.org/doi/10.1021/acs.jmedchem.5c00773
Leave your comments under my linkedIn post here.