Group-Transfer Chimeras for Induced Proximity (GRIPs)
Induced proximity is more than ubiquitination and targeted protein degradation.
In the recent literature we have seen proximity-inducing modalities for phosphorylation and dephosphorylation, deubiquitination, glycosylation and more.
𝗖𝘂𝗿𝗿𝗲𝗻𝘁 𝗹𝗶𝗺𝗶𝘁𝗮𝘁𝗶𝗼𝗻
However, when we induce proximity between our protein of interest (POI) and an effector protein, it can be counterproductive to use inhibitory ligands for recruitment of the effector, as we need to keep it active to do its job.
This limits us to non-inhibitory binders, which exist for less than 1% of all writers/erasers.
Unless we take advantage of the newly developed strategy introducing GRIPs! 💡
𝗞𝗲𝘆 𝗶𝗻𝗻𝗼𝘃𝗮𝘁𝗶𝗼𝗻
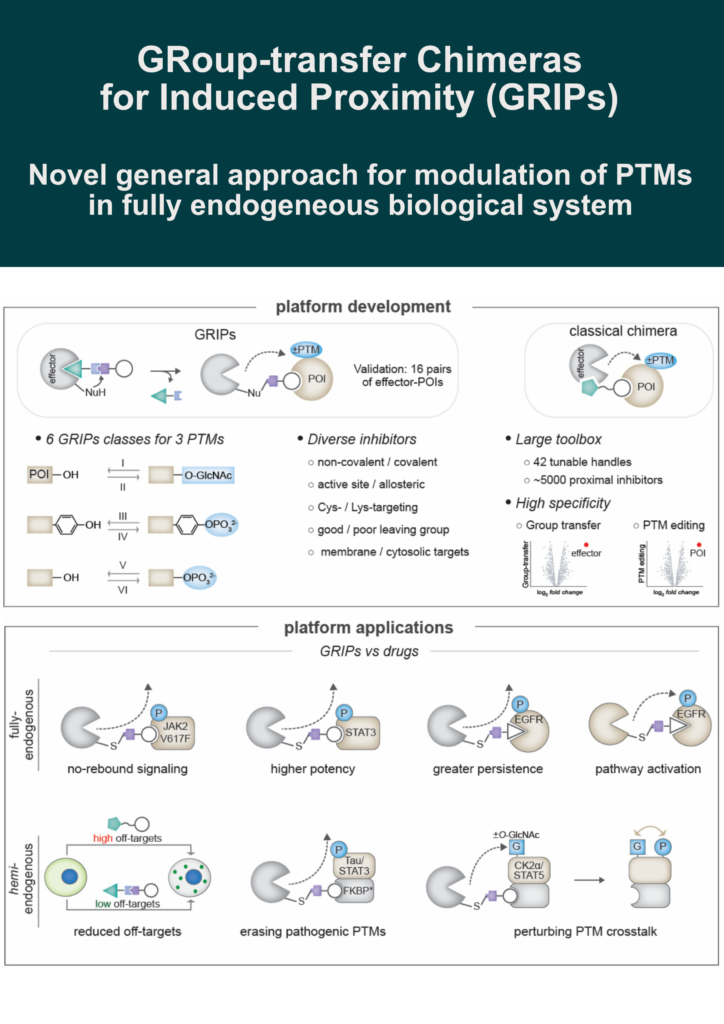
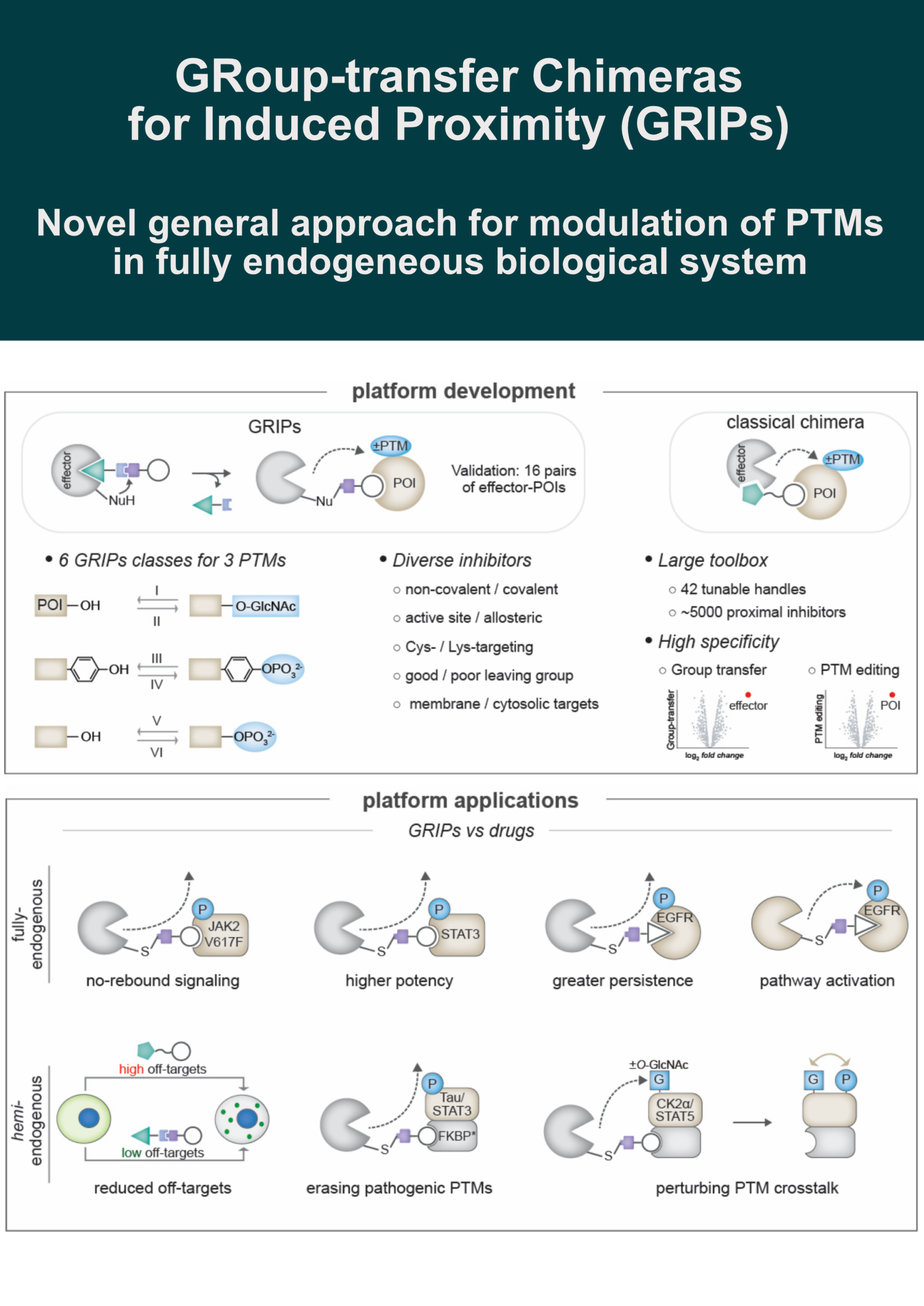
This strategy converts enzyme inhibitors into proximity-inducing chimeras via a cleavable group-transfer handle that covalently appends a POI binder onto a Cys or Lys on the effector, releasing the inhibitor from the active site.
𝗣𝗹𝗮𝘁𝗳𝗼𝗿𝗺 𝘀𝗰𝗼𝗽𝗲
✅ 42 group-transfer handles with distinct reactivity targeting Cys and Lys
✅ ~5,000 plausible inhibitor-residue pairs from PDB mining and chemoproteomics
✅ 6 GRIPs classes: writers or erasers of 3 PTMs (O-GlcNAc, Ser/Thr and Tyr phosphorylation)
✅ 16 effector-POI pairs validated
✅ 4 fully endogenous systems
✅ Compatible with covalent/non-covalent, active-site/allosteric inhibitors
𝗧𝗿𝗮𝗻𝘀𝗹𝗮𝘁𝗲𝘀 𝗶𝗻𝘁𝗼 𝗽𝗵𝗮𝗿𝗺𝗮𝗰𝗼𝗹𝗼𝗴𝗶𝗰𝗮𝗹 𝗲𝗳𝗳𝗲𝗰𝘁𝘀 𝗶𝗻 𝗰𝗲𝗹𝗹𝘂𝗹𝗼
🔸 Baricitinib-based GRIP removed JAK2 rebound signaling in a myeloproliferative neoplasm model (superior to inhibition)
🔸 STAT3-targeting GRIP achieved >4x higher potency than an occupancy-driven inhibitor
🔸 Gefitinib-based GRIPs provided more persistent EGFR inhibition vs. an occupancy-based control
🔸 Osimertinib-derived GRIPs activated EGFR, mimicking EGF, and selectively killed KRAS-mutant cells while sparing wild-type
🔸 AKT-recruiting GRIPs induced Liprin condensate formation with fewer off-targets vs. AMPK-activator chimera
𝗖𝗵𝗲𝗺𝗶𝘀𝘁𝗿𝘆 𝗶𝗻𝘀𝗶𝗴𝗵𝘁𝘀
🔸 Since the reactive warhead undergoes 1,4-addition/elimination sequence, the authors tuned its reactivity by modulating the quality of the leaving group (N to O/S swap, electron-withdrawing groups), which resulted in 6-100x increased reactivity.
🔸 For lysine, they developed SuFA (electrophilic functional group) with ~5x improved stability over NASA. Global proteomics confirmed high specificity of both group transfer and PTM editing.
𝗠𝘆 𝗵𝗶𝗴𝗵𝗹𝗶𝗴𝗵𝘁
What I would like to highlight is that this strategy allows covalent modification of proteins in fully endogenous biological systems while leveraging vast pool of available small molecule ligands.
🧐 What do you think about this strategy to overcome the scalability bottleneck?
Preprint: https://lnkd.in/gkFQiaBf
Leave your comment under my LinkedIn post here.